Introduction
You can determine the equilibrium dissociation constant of an unlabelled ligand by measuring its competition for radioligand binding.
Step by step
Create an XY data table. Enter the logarithm of the concentration of the unlabeled compound into X, and binding into Y. If you have several experimental conditions, place the first into column A, the second into column B, etc. Use subcolumns to enter replicates.
From the data table, click Analyze, choose nonlinear regression, choose the panel of Competition Binding equations, and choose One site - Fit logEC50.
Model
Y=Bottom + (Top-Bottom)/(1+10^(X-LogIC50))
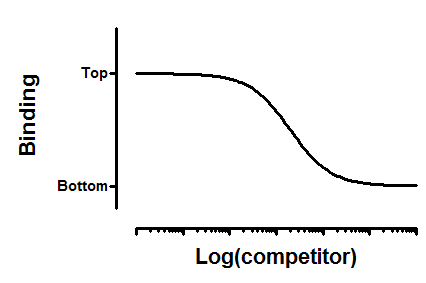
Interpret the parameters
Top and Bottom are plateaus in the units of Y axis.
logIC50 is the log of the concentration of competitor that results in binding half-way between Bottom and Top
Notes
This model is the same as an inhibitory dose-response curve. It fits the logIC50, which is not the same as the Ki of the unlabelled ligand for binding. The Ki depends on the IC50, the concentration of radioligand, and its Kd for binding. You can fit the Ki directly using a different equation.
The analysis assumes that you have one site, and that the binding is reversible and at equilibrium.